|
Growing amounts of genomic data from mixed DNA samples recovered from clinical, animal or environmental samples often result in assemblies and variant determinations with various levels of confidence. cayetanensis due to the complexity of obtaining high quality genomic information from NGS datasets. Unlike the case with foodborne bacteria ( ), the impact of genomics on the development of molecular epidemiological methods based on genomics are not yet fully realized for C. A few PCR targets amplified from geographically distinct strains have been tested for subtyping. cayetanensis organelles-apicoplast and mitochondrion, and whole genome sequences that provide a glimpse into its biology. 
Our group () and others have recently published genomes for the C. cayetanensis is growing over the past 4 years, however, with no immediate solution to the subtyping problem. With the advent of new sequencing methods, the number of genome sequences from C. The lack of animal models or cell culture systems for Cyclospora and the limited availability of its oocysts have hampered its genomics and the development of efficient genotyping tools. Due to the globalization of the food supply, this apicomplexan parasite is prevalent in both endemic regions producing food and non-endemic areas where food is imported . cayetanensis strains during foodborne outbreak investigations.Ĭyclospora cayetanensis is an important apicomplexan parasite causing cyclosporiasis, a common foodborne illness worldwide. A consolidated workflow, presented here, to generate new mitochondrion genomes using our reference-guided de novo assembly approach could be useful in facilitating the generation of other mitochondrion sequences, and in their application for subtyping C. Nucleotide variants were identified in the new and other publicly available genomes in comparison with the mitochondrial reference genome. Using this reference genome, three new mitochondrial genome assemblies were built starting with metagenomic reads generated by sequencing DNA extracted from oocysts present in stool samples from cyclosporiasis patients. In this work, we determined the quality of a high resolution, curated, public mitochondrial genome assembly to be used as a reference genome by applying bioinformatic analyses. The absence of reference genomes to identify nucleotide and structural variants with a high degree of confidence has limited the application of using sequencing data for source tracking during outbreak investigations. To address this issue, development of new methods based on potential genomically-derived markers for strain-level identification has been a priority for the food safety research community. Linking samples from cyclosporiasis patients during foodborne outbreaks with suspected contaminated food sources, using conventional epidemiological methods, has been a persistent challenge. I would be able to speak on the phone if that makes it easier to assist me, and I appreciate and help and advice you can provide.Cyclospora cayetanensis is a coccidian parasite associated with large and complex foodborne outbreaks worldwide. 
Which tool should I use for the trimming? Once it is trimmed can I go straight to mapping, and should I use tophat or bowtie2? My question is how should I combine the data and process it in such a way that it is ready to be mapped?įrom what I've read I will change the data type to fastqsanger, concatenate the technical replicates for the forward and reverse reads and then trim the files. I believe that the files are in fastqsanger format. I therefore have 16 files per experimental group (forward and reverse for each of the 4 biological replicates, each run in two lanes. Each sample was also run in two separate lanes (we were told these technical replicates willl give us more information on isoform expression). 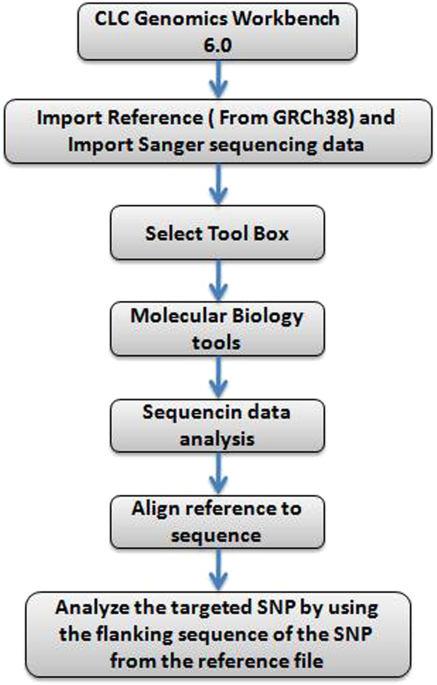
We have 4 biological replicates from 4 experimental groups. They prepared the libraries and performed paired end sequencing. I just got back RNA seq data produced on the HiSeq 4000.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
